There is a tiny device (first invented almost two decades ago) that lets you watch a single cell divide hundreds of times, under tightly-controlled conditions, and yet I almost never meet people who actually use it. This is a shame because there’s many interesting ideas that I think could be uniquely tested in such a device.
Most experiments in biology are instead done in “bulk,” which is not a good way to deeply understand an organism. RNA-seq experiments, for example, are often done by growing lots of cells in a flask, exposing them to some chemical, and then killing all the cells, extracting their RNAs, and sequencing all of them together. The end result of this is just an average, and it completely obscures the messier (and more truthful) stochastic nature of life.
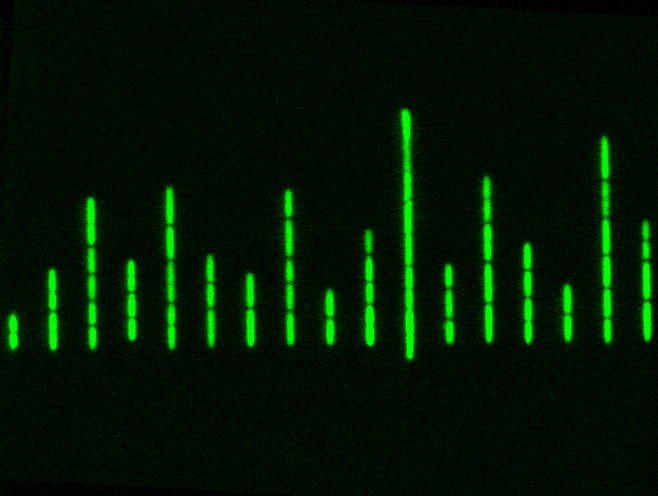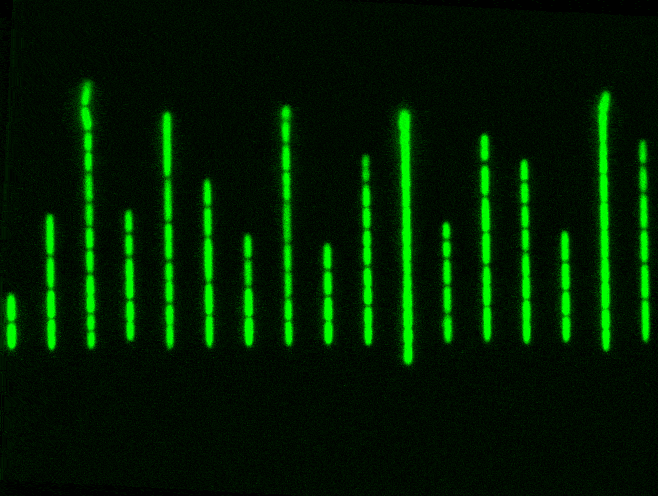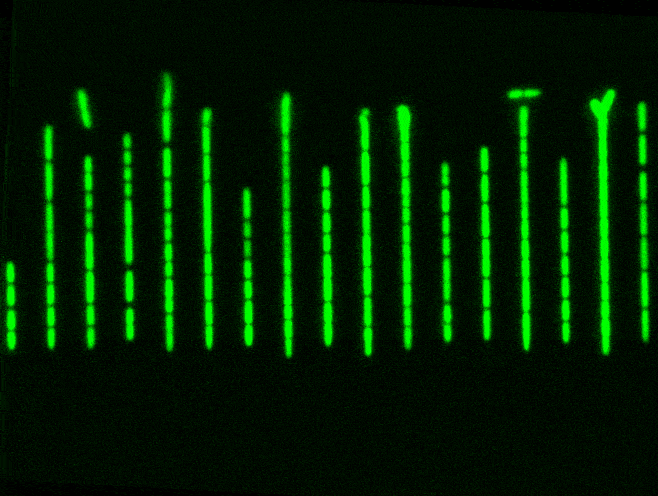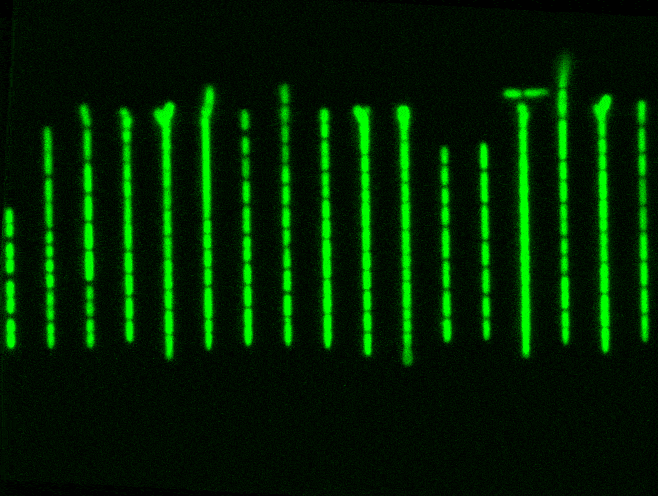
But a Mother Machine, and other microfluidics tools like it, helps to solve this problem. It’s just a tiny device with a long trench through which nutrients flow. Cells travel down this trench and fall into little wells, etched perpendicularly to the main trench. Each of these wells is barely wide enough for a bacterium to fall inside. When a cell falls into a well, it keeps dividing and also has access to fresh nutrients, which are constantly pumped through. Waste molecules are continuously flushed out. As one cell divides into two, then four, then eight, and so on, some cells eventually extend out of the well entirely and get swept away with the current. The cell at the bottom, though, stays put and will keep dividing.
Like many other “great” inventions, the Mother Machine was designed to answer a specific question. In a 2005 paper, some scientists claimed that, when a cell divides, whichever offspring inherits the “old pole” from the mother divides about 2 percent slower with each passing generation. (Said another way: When an E. coli cell divides, it builds a wall down the middle and cuts itself in two at that point. Each “daughter” cell has two ends; one end is made from the wall, and the other end is “old.” This old end gets passed down through the generations, again and again. The authors of the 2005 paper claimed that this constitutes a form of cellular aging, and that cells which inherit the old end are basically less fit than the other daughter cell. Provocative claim!)
The Mother Machine was invented by a small team at Harvard to disprove this hypothesis. In the 2010 paper describing the device, they basically just strapped a camera to the microfluidics chip and recorded the growth rate for tens of thousands of cells, under constant nutrient conditions, for “hundreds of generations.” After tallying all the data, they concluded that “E. coli, unlike all other aging model systems studied to date, has a robust mechanism of growth that is decoupled from cell death.” In other words, growth does not slow down with age, and the 2005 claims were wrong.
Other groups have since made modifications to the original device, using it to revisit classic experiments in molecular biology. In 2018, for example, a Swiss team modified the microfluidic chip to have two input channels, rather than just the one. With two ports, they could expose cells to different growth media at the same time. Or they could switch back-and-forth between the two conditions, or even expose cells to gradients of those conditions.
Now, it has been known since the 1960s that E. coli cells prefer to eat glucose over lactose. When glucose runs out and only lactose is around, the cells activate their lac operon and begin making enzymes to digest it. Jacques Monod and François Jacob shared a Nobel Prize, in 1965, for figuring this out. But nobody had ever actually watched this “switch” at the level of single cells, under tightly-controlled conditions.
But then the Swiss team made their modified Mother Machine. They flooded E. coli cells into the device, trapped them in wells, and switched the inputs between glucose and lactose every four hours. And what they found is that, when lactose comes in to replace glucose, every cell stops growing within three minutes. This outcome is extremely uniform! But the reverse — or the time it takes each cell to switch on its lac operon — is extremely variable. About one-fourth of cells start growing within 25-45 minutes, two-thirds start growing in one-to-three hours, and five percent of cells never grow again at all. By accounting for cells individually, in other words, the Mother Machine enabled these researchers to make observations which could never be made at the population-scale.
And yet, Mother Machines still seem relatively rare! The blueprints are freely available online, but making these devices still requires an understanding of photolithography. The wells are only a micron wide, so they can’t be 3D-printed; one has to make a master mold using photomasks, cast PDMS in that mold, and then cure the polymer into that shape. The original specs only work for E. coli, too. If you wanted to study Bacillus or Caulobacter or yeast, you’d have to redesign the channels with different dimensions. A few companies sell Mother Machines, but they seem to be quite small.
If Mother Machines did become widespread, though (maybe even cheap enough to ship in, say, a $100 kit for students) they could be used to run all kinds of interesting experiments.
One idea is to combine a Mother Machine with a hypermutation tool, such that we can watch cells evolve in real-time. In a recent study, British scientists reported a way to do “highly mutagenic continuous evolution” in E. coli. The beauty of their tool is that it only requires two components: an error-prone DNA polymerase, and a replicon carrying a gene of interest. The error-prone polymerase, which introduces about one mutation per 1,000 bases every ten generations, only copies the DNA on the replicon; it doesn’t touch the host genome. One could take a gene encoding antibiotic resistance (against molecule X) and clone it onto the replicon, transform the whole thing into E. coli, and trap the cells inside a Mother Machine. Then, by exposing the cells to increasing levels of antibiotic Y, one could watch in real time as cells mutate their resistance gene and, perhaps, hit upon a solution that confers resistance against both molecules. This would be a way to study how cells evolve resistance autonomously, at the single-cell level.
Another idea is to use Mother Machines to study how perturbations change a cells’ transcriptome in real-time. Felix Horns (previously in Michael Elowitz’s group at Caltech, now at Arc Institute) created an RNA Exporter tool. The gist is that genes encoding virus-like particles are placed into cells and, when these particles get made, they latch onto RNA molecules and physically carry them out of the cell. Cells are effectively engineered to export their own RNA.
My understanding is that RNA Exporters are relatively unbiased, meaning they have a roughly equal chance of grabbing onto any RNA molecule. The molecules that get carried from the cell, then, are representative of the transcriptome as a whole. If cells carrying RNA Exporters were studied in a Mother Machine, it might be possible to perturb them and measure their transcriptional responses in real time — rather than the classical approach of perturbing millions of cells at once in a flask and doing RNA-seq on the entire population to collect average results.
A third idea is to collect single-cell observations to train a predictive model for molecular burden. Any time we engineer an organism to carry new genes, we are forcing it to execute a function it wouldn’t normally do, thus draining resources that would otherwise go toward growth, DNA repair, and so on. Perhaps we could take 100+ plasmids, each carrying a fluorescent protein, and clone all of them into the same strain of E. coli. Then we could study each strain inside a Mother Machine, carefully quantifying growth rates and fluorescence levels, to map out the full distribution of outcomes for a given plasmid. If we did this enough times (hopefully with some kind of automated data pipeline) we could collect a huge dataset. The resulting model could also help bioengineers design constructs that impose less of a burden on living cells.
I’m not entirely sure why we’re not seeing more of these ideas implemented, or why bioengineers still haven’t fully embraced single-cell experiments. Every university should have a microfluidics facility making custom devices, but I’ve only visited a few of them. Most experiments are still done in bulk, using orders-of-magnitude more cells and reagents than microfluidics would require; and usually the results are less representative of ground truth, too!
It’s a shame, because one of the beautiful things about biology is that each cell is unique and lots of molecular phenomena are highly stochastic, following a distribution of outcomes. Biology is fun because it is not deterministic; and that makes it both richer as a field of study but also more complicated as an engineering medium. A Mother Machine, and other tools like it, help us to actually see these distributions, and we ought to embrace them at scale.