Why do cloning tools still suck? This problem seems like a low-hanging fruit for AI to solve.
Today, if a scientist wants to make a new plasmid or DNA sequence, they often go into their freezer, figure out which DNA sequences they have, upload those DNA sequences to Benchling (or another platform), and then must figure out how to “convert” those sequences into what they want. Should I do Golden Gate or Gibson Assembly? What annealing temperature should I use for my primers? And so on.
There are already tools that help with each of these steps, but has anybody “automated” this decision-making? If so, I’m not familiar with them. (A tool called J5 is probably the closest thing, but it won’t recommend the optimal method given a scientist’s existing sequences and primers.) And if the scientist makes even one error in this multi-step design process — like forgetting about an internal restriction site in a gene — they basically waste an entire week of work.
(You might object to this and say, “But DNA synthesis solves this problem; just synthesize the full plasmid directly!” But people have been saying that for decades at this point, and DNA synthesis costs have not fallen in several years. Cloning DNA remains essential.)
What we need is a fully automated, end-to-end cloning design tool that selects the best method based on a library of existing sequences and primers; a tool that recommends the optimal approach based on cost, speed, and so on. “Design tools” for cloning may not seem like a sexy thing to work on, but whoever solves it will marginally improve the lives of many scientists.
With this in mind, I’ve given $1,500 in microgrants, courtesy of Astera Institute, to two people — Jai Padmakumar and Xavier Bower — who have been thinking about this problem. Bower has already built an open-source prototype, called IceCreamClone.
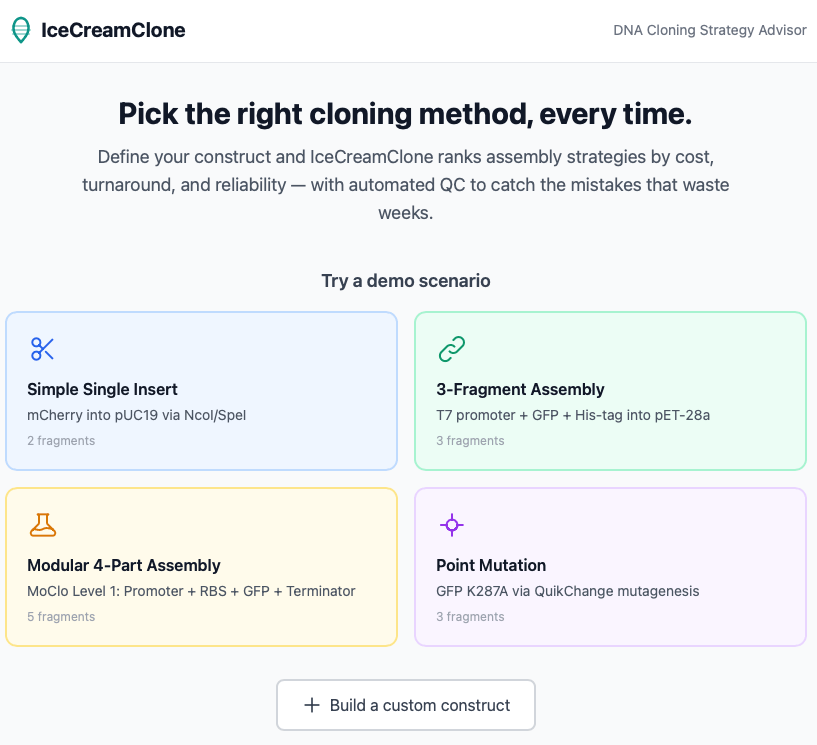
Here’s how a tool like this should work:
First, you specify the plasmid you want to build. Then, you upload your current plasmid library, a collection of DNA sequences already in your inventory, and existing primers. The tool takes these data and outputs multiple cloning protocols based on different metrics, such as lowest cost, fastest speed, or the protocol most likely to be successful. The tool also runs a series of checks on all the sequences to make sure they don’t have internal restriction sites, for example, or weird secondary structures.
It would be particularly cool if scientists using this tool could opt-in to sharing their data. The tool could then prompt them afterwards: How did the cloning go? Can you upload the results? Over time, this feedback data could be used to train predictive models that make cloning far more likely to be successful.
Of course, there are issues with this idea. For one, it requires that people upload their entire catalog of existing sequences + primers, which is quite tedious for some laboratories; especially those with decades of cloning experience. Ideally, these tools would directly integrate with Benchling and Addgene.
Anyway, I continue to think this is a “low-hanging” problem worth working on. Whoever makes an easy-to-use, end-to-end cloning design tool with really good predictive accuracy could presumably make a small business out of it. And, in doing so, you’d make many people happy!